作者:SunCOOL ,写python的农学生。
什么是GFF3格式文件
GFF全称为general feature format,3表示是第三个版本,这种格式主要是用来注释基因组,由tab分割,共9列。详细介绍,可参考这里
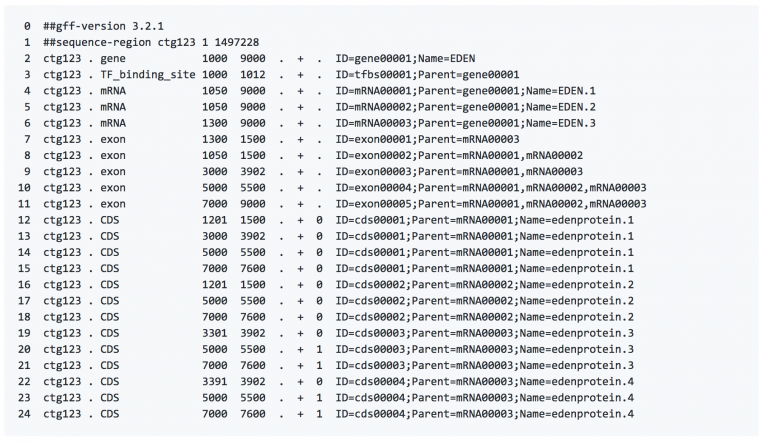
能从GFF3中获取什么
可以获取CDS,EXON,GENE,mRNA再序列中的具体位置,染色体信息,正负链信息
代码
代码的整体流程如下

首先,gff3中,可以根据位置,将其分为一个个的小单元,方便后面的解析,像下面这样
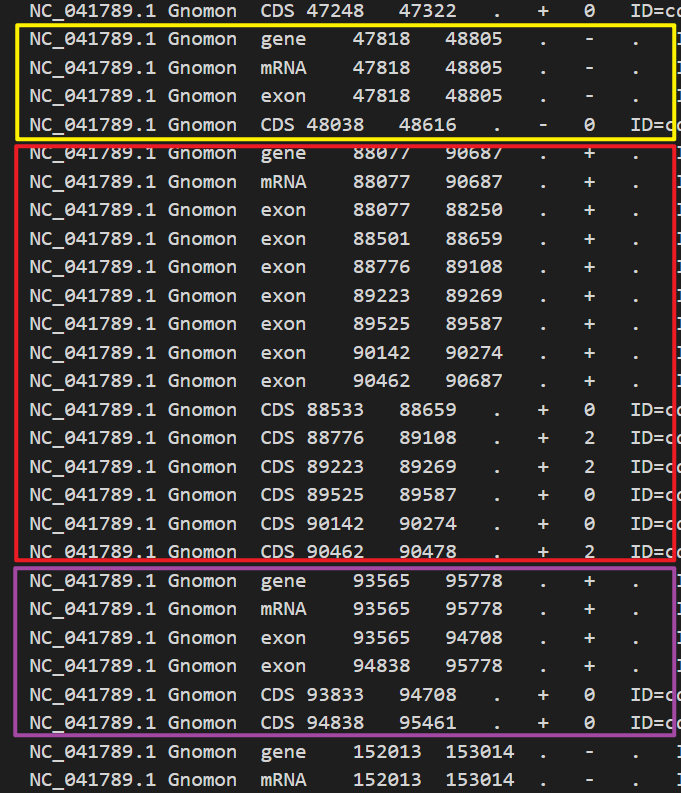
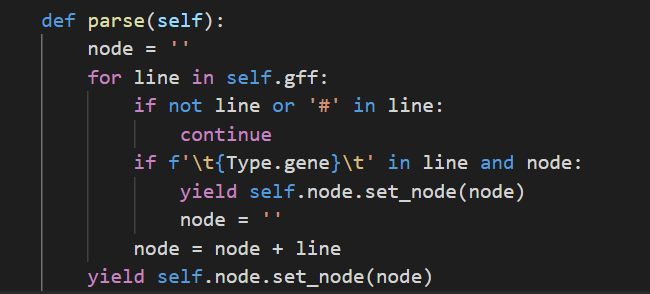
获取了一个个的单元后,接下来就是要对其中ID进行解析提取,方便建立索引,这里使用正则表达式,来进行提取
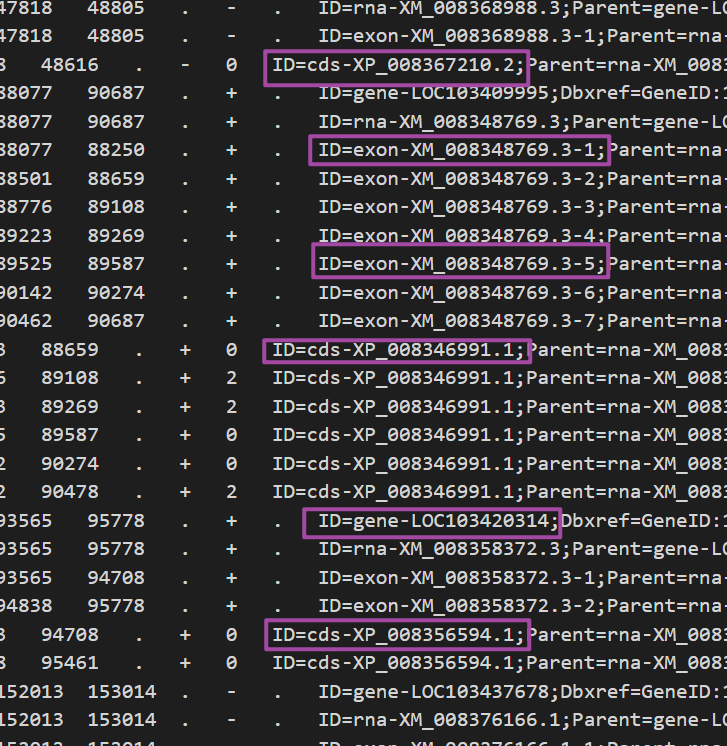
id_patt = '\tID=([\w.-]+);?'
r = re.findall(id_patt, item, re.I | re.S)
在获取了ID以后,由于不同的gff3文件中的ID可能不同,可能会带有一些前缀或者后缀,这里通过添加一系列处理器,来把每一个ID进行二次处理
def create_index(self):
import os
if not os.path.exists(self.index):
conn = sqlite3.connect(self.index)
conn.execute('''CREATE TABLE id_index
(id INT NOT NULL,
id_name TEXT NOT NULL);
''')
c = conn.cursor()
count = 0
insert = 'INSERT INTO id_index(id,id_name) VALUES '
for node in self.parse():
node = self.node_call(node)
sql = ''
for key in node.keys:
sql += '('
sql += f'{count},"{key}"'
sql += '),'
count += 1
sql = insert + sql.strip(',')
c.execute(sql)
conn.commit()
self.gff.seek(0)
self.conn = conn
else:
self.conn = sqlite3.connect(self.index)
建序效果
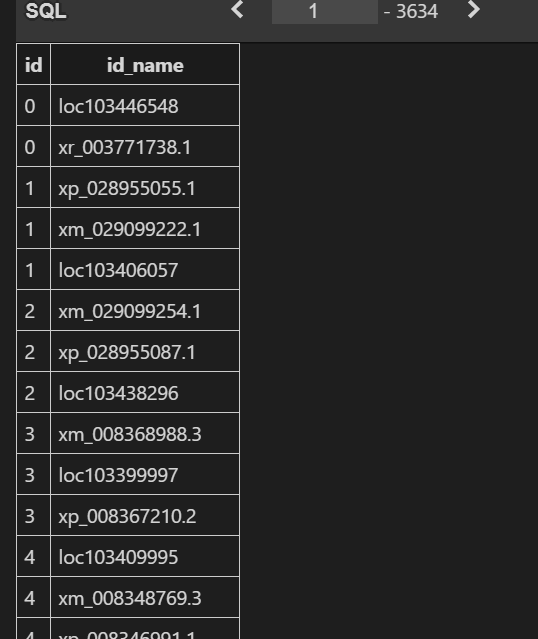
基于gff3文件是由9行构成的,且由tab(\t)分割
def parser_node(self):
node = self.node.split('\n')
uid = str(uuid.uuid4())
for o in node:
o = o.strip()
if not o:
continue
_o = o.split('\t') # 使用\t进行分割
chor = self.safe_lower(_o[0])
type = self.safe_lower(_o[2])
start = self.safe_int(_o[3])
end = self.safe_int(_o[4])
length = end - start + 1
strand = _o[6]
_key = self._get_key(o)
if _key:
key = self.safe_lower(_key[0])
else:
key = 'unknown'
self._nodes.append(dict(zip(
['chro', 'type', 'start', 'end', 'strand', 'key', 'length','uid','o'],
[chor, type, start, end, strand, key, length,uid,o]
)))
查找
通过第一步建序获得的索引文件,可以查得其所对应的GffNode,然后对其解析,然后获取内容。另外,借助python的filter函数,还可以做到筛选功能
def get(self, *, key: str = None, type=None, all=False):
def func(x): return True
if not self._nodes:
self.parser_node()
if all or (key is None and type is None):
func = lambda x:True
elif key and type:
def func(x):
return self.eq(x['key'].lower(), self.safe_lower(key)) and self.eq(x['type'].lower(), type)
elif key or type:
def func(x):
return self.eq(x['key'].lower(), self.safe_lower(key)) or self.eq(x['type'].lower(), type)
r = list(filter(
func,
self._nodes
))
return r
完整代码如下
import re
import sqlite3
import uuid
class Type:
gene = 'gene'
mrna = 'mrna'
cds = 'cds'
lnc_rna = 'lnc_rna'
exon = 'exon'
region = 'region'
pseudogene = 'pseudogene'
transcript = 'transcript'
def to_lower(x):
return str(x).lower()
class GffNode:
id_patt = '\tID=([\w.-]+);?'
key_handles = [to_lower,] # 默认添加一个小写的处理器
def set_node(self, node: str):
self.node = node
self.keys = []
self._nodes = []
return self
def set_key_handle(cls, handle):
cls.key_handles.append(handle) # 添加处理器
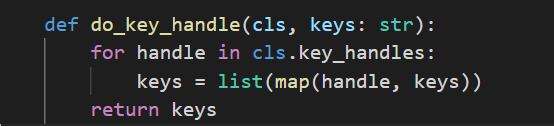
def do_key_handle(cls, keys: str):
for handle in cls.key_handles:
keys = list(map(handle, keys)) # 对提取的id逐个应用处理器
return keys
def _parser_key(self, item: str):
r = re.findall(self.id_patt, item, re.I | re.S) # 提取id
return r or None
def _set_keys(self, keys: str):
if not keys:
return
keys = self.do_key_handle(keys)
self.keys = set(keys) # 获取每一个单元中的所有id并去重
def _get_key(self, item: str):
return self.do_key_handle(self._parser_key(item))
def _call_(self):
keys = self._parser_key(self.node)
self._set_keys(keys)
def eq(self, a, b):
if isinstance(a, str) and isinstance(b, str):
return a in b or b in a or a == b
return False
def __contains__(self, key: str):
key = to_lower(key)
return key in self.keys or any([
self.eq(i, key) for i in self.keys
])
def __len__(self):
return len(self.keys)
def graph(self):
...
def __str__(self):
return f"keys - {len(self)} - {self.keys[0]}"
def parser_node(self):
node = self.node.split('\n')
uid = str(uuid.uuid4()) # 添加uuid,避免出现同一个gene中出现多条mrna难以区分的情况
for o in node:
o = o.strip()
if not o:
continue
_o = o.split('\t')
chor = self.safe_lower(_o[0]) # 染色信息
type = self.safe_lower(_o[2]) # 类型信息
start = self.safe_int(_o[3]) # 开始位置
end = self.safe_int(_o[4]) # 结束位置
length = end - start + 1
strand = _o[6] # 正负链信息
_key = self._get_key(o) # id
if _key:
key = self.safe_lower(_key[0])
else:
key = 'unknown'
self._nodes.append(dict(zip(
['chro', 'type', 'start', 'end', 'strand', 'key', 'length','uid','o'],
[chor, type, start, end, strand, key, length,uid,o] # 保留原始信息o,可以自定义处理
)))
@staticmethod
def safe_lower(o):
if isinstance(o, str):
return o.lower()
return o
@staticmethod
def safe_int(o):
return int(o) # 其实并不安全。。。
def get(self, *, key: str = None, type=None, all=False):
def func(x): return True
if not self._nodes:
self.parser_node()
if all or (key is None and type is None):
func = lambda x:True
elif key and type:
def func(x):
return self.eq(x['key'].lower(), self.safe_lower(key)) and self.eq(x['type'].lower(), type)
elif key or type:
def func(x):
return self.eq(x['key'].lower(), self.safe_lower(key)) or self.eq(x['type'].lower(), type)
r = list(filter(
func,
self._nodes
))
return r
class Gff:
def __init__(self, gff_file, id_file):
self.gff = open(gff_file, 'r', encoding='utf-8')
self.id_file = open(id_file, 'r', encoding='utf-8')
self.index = gff_file + '.index'
self.node = GffNode()
def run(self):
self.create_index() # 建序
self.ids = self.read_ids() # 读取ID文件
def set_key_handle(self, handle):
self.node.set_key_handle(handle)
def create_index(self):
import os
if not os.path.exists(self.index):
conn = sqlite3.connect(self.index)
conn.execute('''CREATE TABLE id_index
(id INT NOT NULL,
id_name TEXT NOT NULL);
''')
c = conn.cursor()
count = 0
insert = 'INSERT INTO id_index(id,id_name) VALUES '
for node in self.parse():
node = self.node_call(node)
sql = ''
for key in node.keys:
sql += '('
sql += f'{count},"{key}"'
sql += '),'
count += 1
sql = insert + sql.strip(',')
c.execute(sql)
conn.commit()
self.gff.seek(0)
self.conn = conn
else:
self.conn = sqlite3.connect(self.index)
@staticmethod
def node_call(node):
node._call_()
return node
def parse(self):
node = ''
for line in self.gff:
if not line or '#' in line:
continue
if f'\t{Type.gene}\t' in line and node:
yield self.node.set_node(node)
node = ''
node = node + line
yield self.node.set_node(node)
def read_ids(self):
ids = self.id_file.readlines()
ids = [i.replace('\n', '').lower() for i in ids]
conn = self.conn
_ids = []
for id in ids:
_search_sql = 'select id,id_name from id_index where id_name like "%s"' % id
rows = conn.execute(_search_sql)
rows = list(rows)
if rows:
_ids.append(rows[0])
_ids.sort(key=lambda x: x[0], reverse=True)
return _ids
def search(self, *, key=None, type=None, all=False, use_strict=True):
"""
key 即ID
type 是gene、exon、cds等
all 返回整个单元的所有结构信息,优先级最高(此时key或者type会失效)
use_strict 返回结果中是否要通过key在做一次筛选,一般用不到。在多顺反子中可能会用的到。
"""
count = 0
ids = []
keys = []
if self.ids:
for _id in self.ids:
if _id[0] not in ids:
ids.append(_id[0])
keys.append(_id[1])
else:
return []
first = ids.pop()
data = []
for node in self.parse():
if count == first:
node = self.node_call(node)
data = data + node.get(key=key, type=type, all=all)
if not ids:
break
first = ids.pop()
count += 1
return list(filter(
lambda x: x['key'] in keys if use_strict else True,
data
))
if __name__ == "__main__":
p = 'seq\genomic.gff3'
id_file = 'hmm\ids.txt' # 每一行放置一个id
p = Gff(p, id_file)
def func(x):
if 'gene' in x:
return x.split('-')[1]
if 'cds' in x:
return x.split('-', 1)[1]
if 'exon' in x:
return x.split('-')[1]
if 'rna' in x:
return x.split('-', 1)[1]
return x
p.set_key_handle(func) # 添加自定义的ID处理器
# 添加处理器的目的是为了将gff3中的id转换成你的id_file中的id
# 所以可以添加多个处理器
# 一步步的进行处理,直到id和id_file中的相同
p.run()
d = p.search(type=Type.gene, use_strict=False)
import json
print(json.dump(d, open('gene.json', 'w'))) # 写入json文件
赞赏作者

Python中文社区作为一个去中心化的全球技术社区,以成为全球20万Python中文开发者的精神部落为愿景,目前覆盖各大主流媒体和协作平台,与阿里、腾讯、百度、微软、亚马逊、开源中国、CSDN等业界知名公司和技术社区建立了广泛的联系,拥有来自十多个国家和地区数万名登记会员,会员来自以工信部、清华大学、北京大学、北京邮电大学、中国人民银行、中科院、中金、华为、BAT、谷歌、微软等为代表的政府机关、科研单位、金融机构以及海内外知名公司,全平台近20万开发者关注。
长按扫码添加“Python小助手”

▼点击成为社区会员 喜欢就点个在看吧





